The author is associate professor of dairy cattle genetics at Penn State University.
Genomic evaluations were officially released for the first time in January of 2009. The third anniversary gives us a good opportunity to review the accuracy of those initial predictions.
There were more than 2,000 Holstein bulls in the initial genomic release with no daughters. Of those, 354 had at least 100 daughters by August of 2010. There is some importance to the 100-daughter benchmark. Requiring 100 daughters in August of 2010 provides time for cows to be culled and evaluate their herd life more accurately today.
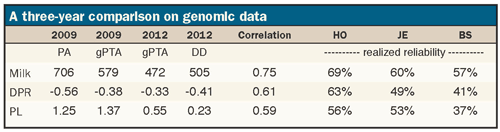
Some statistics for those bulls are summarized in the table for milk yield, daughter pregnancy rate (DPR) and productive life (PL). We could look at many traits, but these will provide a good sense of how genomic predictions have worked without overwhelming us with numbers.
A side-by-side comparison
Included in the table are the average genomic predicted transmitting abilities (gPTA) and parent averages (PA) for each trait in 2009. There was a genetic base change in 2010, so I've converted the 2009 averages to the current base. We will compare the numbers to December 2012 gPTA and daughter deviations (DD).
Comparisons with the DD are the most ideal because they are based only on daughter records, whereas 2012 gPTA includes a mix of daughter records, pedigree, and genetic marker information. As a bull accumulates daughters, the pedigree and genomic information is weighted less and the gPTA will eventually become nearly identical to DD.
It is apparent that milk and PL were overestimated when considering only the PA. This has always been a challenge in our progeny testing programs. The gPTA helped to reduce that bias for yield, but it still overestimated milk by 74 pounds and one month for PL when compared to DD.
Perhaps more important is how accurately the gPTA ranked bulls that did not have daughters. The correlations between 2009 gPTA and 2012 DD are also included in the table, and range from 0.59 for PL to 0.75 for yield.
These correlations can be used to derive the realized reliability of the 2009 gPTA. The realized reliabilities were within 3 percent of the published reliabilities for milk and DPR. In other words, the amount of re-ranking among bulls for those traits was similar to what we expected. The numbers suggest the genomic PL evaluations were not as accurate as we had hoped.
Quality control continues
The scientists at AIPL responsible for genomic evaluations perform similar quality control tests for all traits in the Holstein breed, and in the Jerseys and Brown Swiss, as well. The traits that still give some troubles in Holsteins are PL, sire calving ease, and sire stillbirth. Type traits evaluations have generally performed well for Holsteins.
I've used AIPL's test data from this past fall to also show you the realized reliabilities for Jerseys and Brown Swiss. The results for PL and DPR are in line with expectations for Jersey, but slightly less than expected for yield traits. The results for Brown Swiss are not significantly different than expected, but the relatively lower reliabilities highlight the challenge that is faced by our breeds with smaller populations. It takes a lot of genomically tested sires that have many daughters to generate robust gPTA.
All breeds will benefit from more testing of males and females, including more animals from the bottom half of the populations so that we have something to compare with the superior cows and bulls. These early results also reinforce how critical it is that we continue to invest in our progeny testing system, particularly for lower heritability and calving traits.
Not a direct comparison
The numbers suggest that we can make reasonable comparisons among genomically tested young sires for most traits, which was not true before we had genomic predictions. However, the declines from 2009 to 2012 do suggest that we have to be careful when comparing progeny tested to nonprogeny tested bulls.
Another way of looking at this is to compare the merit of top progeny tested and genomic young sires in 2009 to their merit now that they all have daughters. The top 25 bulls with no daughters in 2009 had an average Net Merit of $620 (on today's genetic base) compared with $495 for the 25 highest first crop daughter bulls that year. In 2012, those two sets of bulls had an identical average of $479. In other words, the bulls with no daughters were not as superior as initially believed when compared to progeny tested bulls. That said, we have made some improvements.
There was a somewhat controversial rescaling of cow gPTA in April of 2010 in order to reduce the tendency to overestimate gPTA for nonprogeny tested bulls. That change should help move us toward being able to make more direct comparisons between bulls with and without daughters.
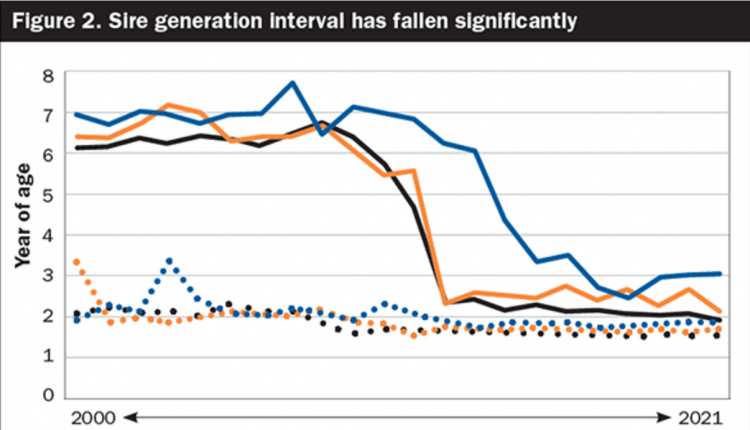
The top 25 Net Merit sires that had no daughters in August of 2010 but now have at least 50 milking daughters experienced a much smaller decline in Net Merit ($64 versus $125 in the 2009 group) and have a slight advantage over the first-crop bulls from that time. We can probably compare progeny tested to non-progeny tested more directly now than in 2009, but that is a subject that we will need to revisit down the road to have confidence that those comparisons are valid. It is also important to note that we are only now just beginning to see the young bulls with greatly reduced generation intervals become available, so the real advances from genomics are yet to be realized.
Not complete agreement
It is my sense that people's perception of genomics takes on a different perspective based on whether you are a population geneticist (like me) and or a producer-breeder. Some loss of accuracy is okay from the population geneticist standpoint because it will be more than offset by a reduction in generation interval. If a bull's proof turns out to have been inflated, there will be another bull that was correct to become the next great sire-of-sons and the population will advance rapidly.
That may be okay, but it wasn't my cow that was bred to the bull that wasn't better in calving ease after all or my cow that was flushed to a suddenly mediocre bull. That is a difference in perspective that genomics won't eliminate but is a difference we can acknowledge and appreciate.
There is no question that the scientists involved with genomic predictions have done a tremendous job and are constantly looking for ways to improve the system. I have a lot of confidence that our breeders will use these tools to continue their development of exceptional cattle. For those breeders using genomic young sires, it is still advisable to select a variety of bulls in order to spread your risk.
Genomic evaluations were officially released for the first time in January of 2009. The third anniversary gives us a good opportunity to review the accuracy of those initial predictions.
There were more than 2,000 Holstein bulls in the initial genomic release with no daughters. Of those, 354 had at least 100 daughters by August of 2010. There is some importance to the 100-daughter benchmark. Requiring 100 daughters in August of 2010 provides time for cows to be culled and evaluate their herd life more accurately today.
Some statistics for those bulls are summarized in the table for milk yield, daughter pregnancy rate (DPR) and productive life (PL). We could look at many traits, but these will provide a good sense of how genomic predictions have worked without overwhelming us with numbers.
A side-by-side comparison
Included in the table are the average genomic predicted transmitting abilities (gPTA) and parent averages (PA) for each trait in 2009. There was a genetic base change in 2010, so I've converted the 2009 averages to the current base. We will compare the numbers to December 2012 gPTA and daughter deviations (DD).
Comparisons with the DD are the most ideal because they are based only on daughter records, whereas 2012 gPTA includes a mix of daughter records, pedigree, and genetic marker information. As a bull accumulates daughters, the pedigree and genomic information is weighted less and the gPTA will eventually become nearly identical to DD.
It is apparent that milk and PL were overestimated when considering only the PA. This has always been a challenge in our progeny testing programs. The gPTA helped to reduce that bias for yield, but it still overestimated milk by 74 pounds and one month for PL when compared to DD.
Perhaps more important is how accurately the gPTA ranked bulls that did not have daughters. The correlations between 2009 gPTA and 2012 DD are also included in the table, and range from 0.59 for PL to 0.75 for yield.
These correlations can be used to derive the realized reliability of the 2009 gPTA. The realized reliabilities were within 3 percent of the published reliabilities for milk and DPR. In other words, the amount of re-ranking among bulls for those traits was similar to what we expected. The numbers suggest the genomic PL evaluations were not as accurate as we had hoped.
Quality control continues
The scientists at AIPL responsible for genomic evaluations perform similar quality control tests for all traits in the Holstein breed, and in the Jerseys and Brown Swiss, as well. The traits that still give some troubles in Holsteins are PL, sire calving ease, and sire stillbirth. Type traits evaluations have generally performed well for Holsteins.
I've used AIPL's test data from this past fall to also show you the realized reliabilities for Jerseys and Brown Swiss. The results for PL and DPR are in line with expectations for Jersey, but slightly less than expected for yield traits. The results for Brown Swiss are not significantly different than expected, but the relatively lower reliabilities highlight the challenge that is faced by our breeds with smaller populations. It takes a lot of genomically tested sires that have many daughters to generate robust gPTA.
All breeds will benefit from more testing of males and females, including more animals from the bottom half of the populations so that we have something to compare with the superior cows and bulls. These early results also reinforce how critical it is that we continue to invest in our progeny testing system, particularly for lower heritability and calving traits.
Not a direct comparison
The numbers suggest that we can make reasonable comparisons among genomically tested young sires for most traits, which was not true before we had genomic predictions. However, the declines from 2009 to 2012 do suggest that we have to be careful when comparing progeny tested to nonprogeny tested bulls.
Another way of looking at this is to compare the merit of top progeny tested and genomic young sires in 2009 to their merit now that they all have daughters. The top 25 bulls with no daughters in 2009 had an average Net Merit of $620 (on today's genetic base) compared with $495 for the 25 highest first crop daughter bulls that year. In 2012, those two sets of bulls had an identical average of $479. In other words, the bulls with no daughters were not as superior as initially believed when compared to progeny tested bulls. That said, we have made some improvements.
There was a somewhat controversial rescaling of cow gPTA in April of 2010 in order to reduce the tendency to overestimate gPTA for nonprogeny tested bulls. That change should help move us toward being able to make more direct comparisons between bulls with and without daughters.
The top 25 Net Merit sires that had no daughters in August of 2010 but now have at least 50 milking daughters experienced a much smaller decline in Net Merit ($64 versus $125 in the 2009 group) and have a slight advantage over the first-crop bulls from that time. We can probably compare progeny tested to non-progeny tested more directly now than in 2009, but that is a subject that we will need to revisit down the road to have confidence that those comparisons are valid. It is also important to note that we are only now just beginning to see the young bulls with greatly reduced generation intervals become available, so the real advances from genomics are yet to be realized.

Not complete agreement
It is my sense that people's perception of genomics takes on a different perspective based on whether you are a population geneticist (like me) and or a producer-breeder. Some loss of accuracy is okay from the population geneticist standpoint because it will be more than offset by a reduction in generation interval. If a bull's proof turns out to have been inflated, there will be another bull that was correct to become the next great sire-of-sons and the population will advance rapidly.
That may be okay, but it wasn't my cow that was bred to the bull that wasn't better in calving ease after all or my cow that was flushed to a suddenly mediocre bull. That is a difference in perspective that genomics won't eliminate but is a difference we can acknowledge and appreciate.
There is no question that the scientists involved with genomic predictions have done a tremendous job and are constantly looking for ways to improve the system. I have a lot of confidence that our breeders will use these tools to continue their development of exceptional cattle. For those breeders using genomic young sires, it is still advisable to select a variety of bulls in order to spread your risk.
This article appears on page 303 of the April 25, 2012 issue of Hoard's Dairyman.